Citric acid cross-linking is an effective strategy for modifying citrus pectin to enhance its performance in sustainable packaging applications. In this article, Chandra Mohan Chandrasekar, Daniele Carullo, Francesco Saitta, Tommaso Bellesia, Elena Caneva, Chiara Baschieri, Marco Signorelli, Dimitrios Fessas, Stefano Farris and Davide Romano, investigated the structural changes induced by citric acid cross-linking and their influence on the properties of nanocellulose-reinforced packaging films..
The authors demonstrated that cross-linking significantly alters the structure–property relationship of the biopolymer matrix, leading to improved film integrity and modified surface morphology. These results provide valuable insight into biopolymer modification strategies for the development of environmentally friendly packaging materials.
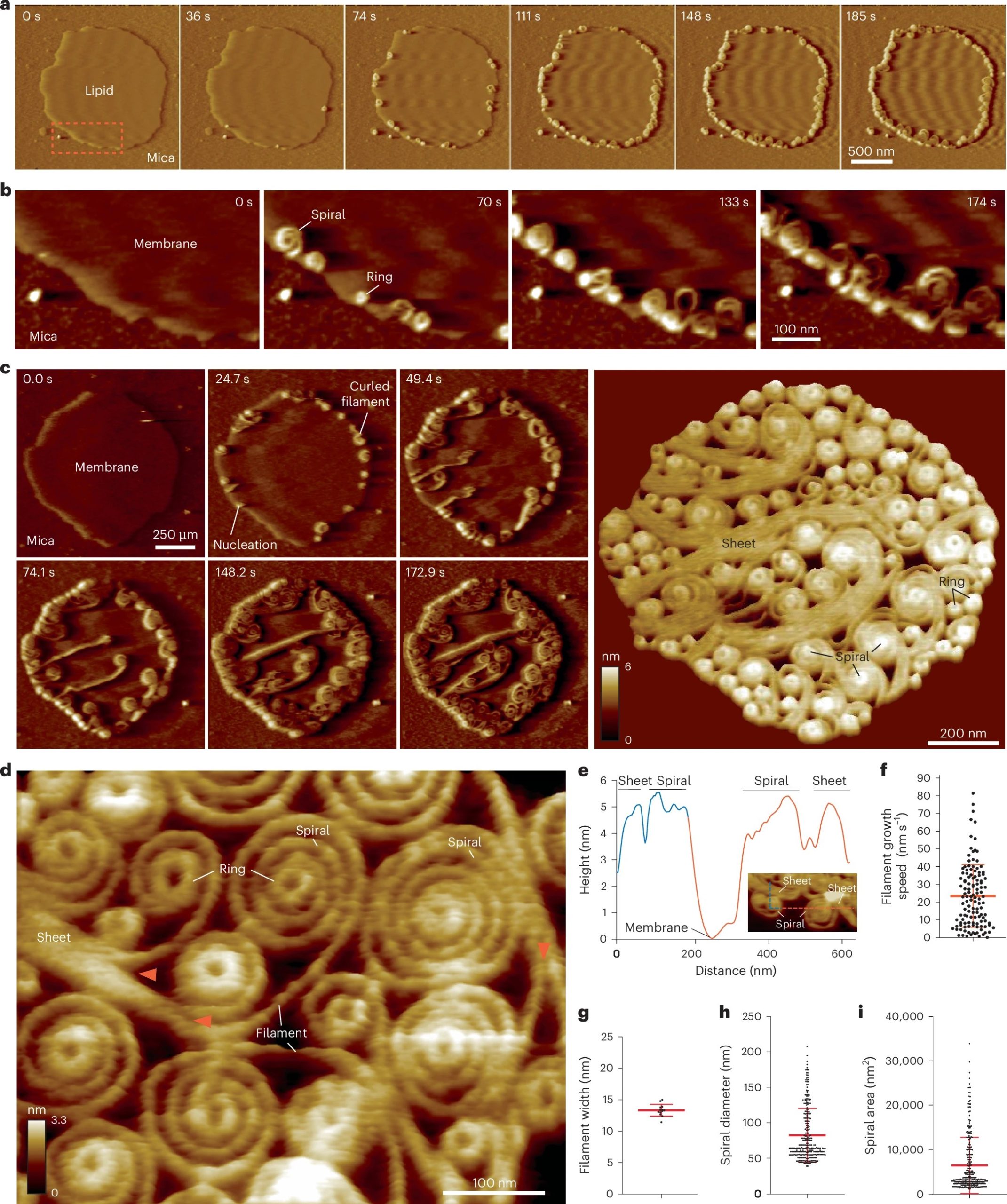
Atomic force microscopy (AFM) was employed to characterize the surface morphology of the films. Measurements were performed using a commercially available AFM instrument operated in contact resonance amplitude imaging (CRAI) mode. A NanoWorld Arrow-FMR AFM probe was used.
This AFM probe features a rectangular beam with a triangular free end and a tetrahedral tip (tip radius ~10 nm, tip height 10–15 μm), with a spring constant of 2.8 N/m and a resonance frequency of 75 kHz. Root mean square surface roughness values were calculated from at least five height-mode images.
Full citation: Chandrasekar, C. M.; Carullo, D.; Saitta, F.; Bellesia, T.; Caneva, E.; Baschieri, C.; Signorelli, M.; Fessas, D.; Farris, S.; Romano, D. “Structural elucidation of citric acid cross-linked pectin and its impact on the properties of nanocellulose-reinforced packaging films.” International Journal of Biological Macromolecules 2025, 333(2), 148869. https://doi.org/10.1016/j.ijbiomac.2025.148869
Attribution 4.0 International By exercising the Licensed Rights (defined below), You accept and agree to be bound by the terms and conditions of this Creative Commons Attribution 4.0 International Public License (“Public License”). To the extent this Public License may be interpreted as a contract, You are granted the Licensed Rights in consideration of Your acceptance of these terms and conditions, and the Licensor grants You such rights in consideration of benefits the Licensor receives from making the Licensed Material available under these terms and conditions. https://creativecommons.org/licenses/by/4.0/legalcode.en