Color centers in hexagonal boron nitride (hBN) are promising candidates as quantum light sources for future technologies. *
In the article “Nanoscale resolved mapping of the dipole emission of hBN color centers with a scattering-type scanning near-field optical microscope “, Iris Niehues, Daniel Wigger, Korbinian Kaltenecker, Annika Klein-Hitpass , Philippe Roell, Aleksandra K. Dąbrowska, Katarzyna Ludwiczak, Piotr Tatarczak, Janne O. Becker , Robert Schmidt, Martin Schnell, Johannes Binder, Andrzej Wysmołek and Rainer Hillenbrand utilize a scattering-type near-field optical microscope (s-SNOM) to study the photoluminescence (PL) emission characteristics of such quantum emitters in metalorganic vapor phase epitaxy grown hBN. *
On the one hand, Iris Niehues et al. demonstrate direct near-field optical excitation and emission through interaction with the nanofocus of the AFM tip resulting in a subdiffraction limited tip-enhanced PL hotspot. *
On the other hand, the authors show that indirect excitation and emission via scattering from the AFM tip significantly increases the recorded PL intensity. This demonstrates that the tip-assisted PL (TAPL) process efficiently guides the generated light to the detector. *
Iris Niehues et al. apply the TAPL method to map the in-plane dipole orientations of the hBN color centers on the nanoscale. This work promotes the widely available s-SNOM approach to applications in the quantum domain including characterization and optical control. *
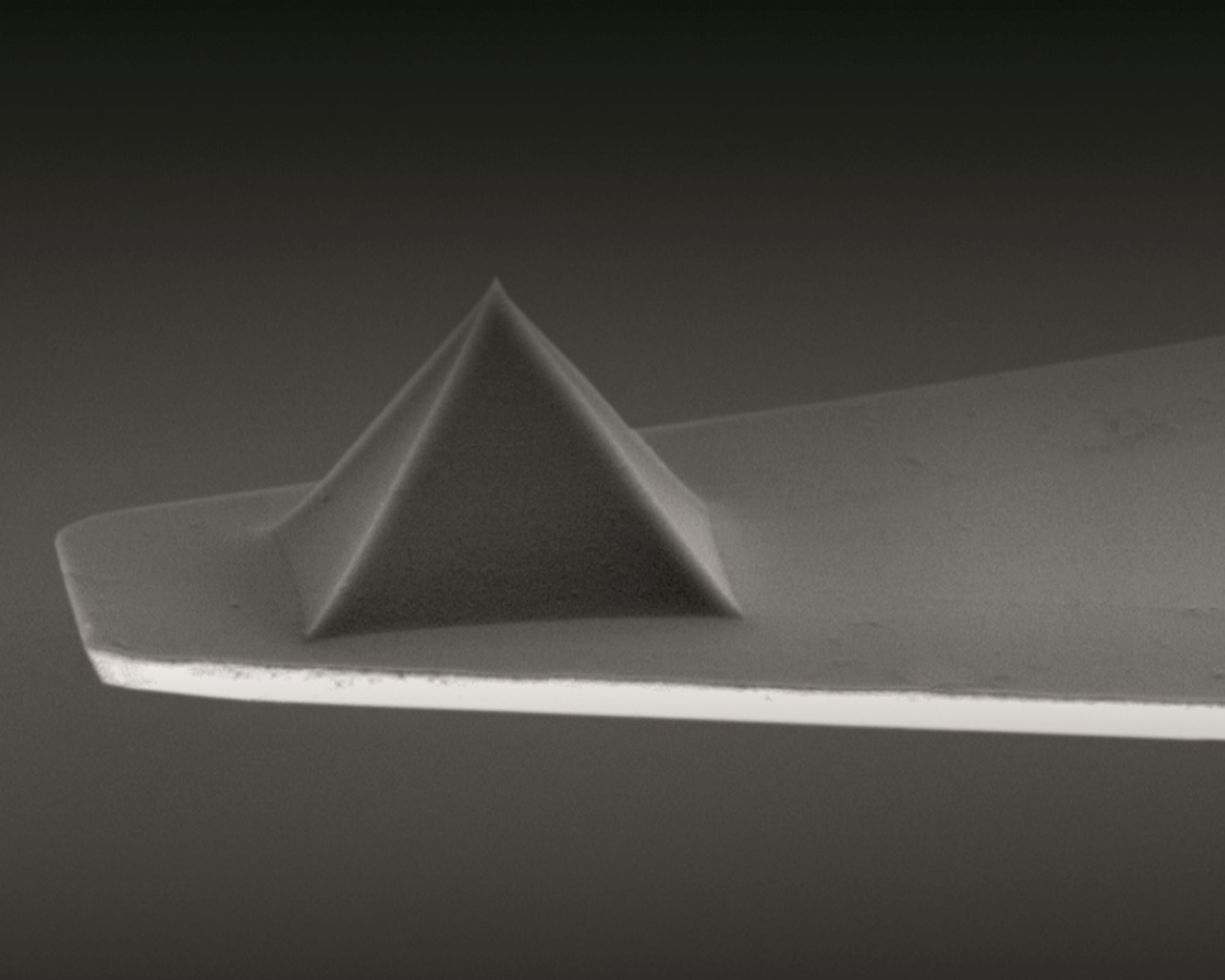
The investigation utilizes a scattering-type near-field optical microscope employing a metallized Arrow AFM tip ( NanoWorld Arrow-NCPt AFM probe) illuminated by monochromatic laser light. *
The AFM tip acts as an optical antenna, transforming the incident p-polarizedlight into a highly focused near field at the AFM tip apex, the so-called nanofocus. *
The nanofocus interacts with the sample leading to modified scattering from the AFM tip and encoding local sample properties.
In conventional s-SNOM operation, the elastically scattered light is recorded as function of sample position (note that the sample is scanned), yielding near-field optical images with a spatial resolution down to 10 nm. *
To supress background scattering, the AFM is operated in tapping mode and the detector signal is demodulated at a higher harmonic of the AFM tip’s oscillation frequency. *
In the article, Iris Niehues et al. use the s-SNOM instrument to study PL from individual hBN color centers. *
To that end, the inelastically tip-scattered light is recorded with a grating spectrometer coupled to a CCD camera. Note that signal demodulation has not been possible with the use of a CCD camera so far. It may be achieved employing a photomultiplier tube or similar. Importantly, the authors’ s-SNOM setup includes a high-quality, silver-protected off-axis parabolic mirror with a numerical aperture (NA) of 0.72, which optimizes the focusing and collection efficiency of the optical system and is crucial for the performed PL measurements. *
Characterization of photoluminescence mapping
In the specific experiments performed by “, Iris Niehues et al., the authors employ the near-field optical microscope in tapping mode, with low oscillation amplitudes between 20 nm and 30 nm, to detect PL signals influenced by the presence of the metallic AFM tip. *
They use standard metallic Arrow AFM tips (NanoWorld Arrow-NCPt) Throughout this study, Iris Niehues et al., use a 532 nm (2.33 eV) laser for the optical excitation of the hBN color centers. *

Photoluminescence (PL) measurement of a single color center taken with an AFM tip. The images are shown with the same color bar for better comparison of the observed PL intensities. (a) PL intensity map without the tip showing a diffraction limited emission spot. (b) PL spectrum of the studied emitter recorded with an extended integration time inside the arc in (c). The zero-phonon line (ZPL) and optical phonon sidebands (PSBs) of 160 meV are marked as well as the broad background PL (black line). (c) PL map of the same emitter with the AFM tip showing two subdiffraction limit features marked as “dot” and “arc.” (d) Lineprofiles along the dashed lines in (a) in black and (c) in red (dark measurement, bright Gaussian fits). The fitted full widths at half maximum (FWHM) are 110 nm (dot), 209 nm (arc), and 1,418 nm (w/o tip). (e) Schematic of the interference between direct and indirect excitation/emission of the color center via the AFM tip (TAPL). Inset shows the nanofocus interaction at the location of the color center explaining the dot (TEPL). (f) Analytical reproduction of the TAPL arc in (c) applying the model in (e).
*Iris Niehues, Daniel Wigger, Korbinian Kaltenecker, Annika Klein-Hitpass , Philippe Roell, Aleksandra K. Dąbrowska, Katarzyna Ludwiczak, Piotr Tatarczak, Janne O. Becker , Robert Schmidt, Martin Schnell, Johannes Binder, Andrzej Wysmołek and Rainer Hillenbrand
Nanoscale resolved mapping of the dipole emission of hBN color centers with a scattering-type scanning near-field optical microscope
Nanophotonics, vol. 14, no. 3, 2025, pp. 335-342
DOI: https://doi.org/10.1515/nanoph-2024-0554
Open Access The article “Nanoscale resolved mapping of the dipole emission of hBN color centers with a scattering-type scanning near-field optical microscope” by Iris Niehues, Daniel Wigger, Korbinian Kaltenecker, Annika Klein-Hitpass , Philippe Roell, Aleksandra K. Dąbrowska, Katarzyna Ludwiczak, Piotr Tatarczak, Janne O. Becker , Robert Schmidt, Martin Schnell, Johannes Binder, Andrzej Wysmołek and Rainer Hillenbrand is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.